Software Engineering Tips and Best Practices for Data Science
- The Tech Platform

- Sep 12, 2020
- 7 min read
With great code comes great machine learning
If you’re into data science you’re probably familiar with this workflow: you start a project by firing up a jupyter notebook, then begin writing your python code, running complex analyses, or even training a model. As the notebook file grows in size with all the functions, the classes, the plots, and the logs, you find yourself with an enormous blob of monolithic code sitting up in one place in front of you. If you’re lucky, things can go well. Good for you then!
However, jupyter notebooks hide some serious pitfalls that may turn your coding into a living hell. Let’s see how this happens and then discuss coding best practices to prevent it
The problems with Jupyter Notebook ⛔️
source: datascience.foundation
Quite often, things may not go the way you intend if you want to take your jupyter prototyping to the next level. Here are some situations I myself encountered while using this tool and that should sound familiar to you:
With all the objects (functions or classes) defined and instantiated in one place, maintainability becomes really hard: even if you want to make a small change to a function, you have to locate it somewhere in the notebook, fix it and rerun the code all over again. You don’t want that, believe me. Wouldn’t be simple to have your logic and processing functions separated in external scripts?
Because of its interactivity and instant feedback, jupyter notebooks push data scientists to declare variables in the global namespace instead of using functions. This is considered bad practice in python development because it limits effective code reuse. It also harms reproducibility because your notebook turns into a large state machine holding all your variables. In this configuration, you’ll have to remember which result is cached and which is not and you’ll also have to expect other users to follow your cell execution order.
The way notebooks are formatted behind the scenes (JSON objects) makes code versioning difficult. This is why I rarely see data scientists using GIT to commit different versions of a notebook or merging branches for specific features. Consequently, team collaboration becomes inefficient and clunky: team members start exchanging code snippets and notebooks via e-mail or Slack, rolling back to a previous version of the code is a nightmare, and the file organization starts to be messy. Here’s what I commonly see in projects after two or three weeks of using a jupyter notebook without proper versioning:
analysis.ipynb
analysis_COPY(1).ipynb
analysis_COPY(2).ipynb
analysis_FINAL.ipynb
analysis_FINAL_2.ipynbJupyter notebooks are good for exploration and quick prototyping. They’re certainly not designed for reusability or production-use. If you developed a data processing pipeline using a jupyter notebook, the best you can state is that your code is only working on your laptop or your VM in a linear synchronous fashion following the execution order of the cells. This doesn’t say anything about the way your code would behave in a more complex environment with, for instance, larger input datasets, other asynchronous parallel tasks, or less allocated resources. Notebooks are in fact hard to test since their behavior is sometimes unpredictable.
As someone who spends most of his time on VSCode taking advantage of powerful extensions for code linting, style formatting, code structuring, autocompletion, and codebase search, I can’t help but feel a bit powerless when switching back to jupyter. Compared to VSCode, jupyter notebook lacks extensions that enforce coding best practices.
Ok folks, enough bashing for now. I honestly love jupyter and I think it’s great for what’s designed to do. You can definitely use it to bootstrap small projects or quickly prototype ideas.
But in order to ship these ideas in an industrial fashion, you have to follow software engineering principles that happen to get lost when data scientists use notebooks. So let’s review some of them together and see why they’re important.
Tips to make your code great again 🚀
*These tips have been compiled from different projects, meetups I attended, and discussions with software engineers and architects I’ve worked with by the past. If you have other suggestions and ideas to share, feel free to bring your contributions in the comment section and I’ll credit your answer in the post.
*The following sections assume that we’re writing python scripts. Not notebooks.
1 —Clean your code 🧼️
Photo by Florian Olivo on Unsplash
One of the most important aspects of code quality is clarity. Clear and readable code is crucial for collaboration and maintainability.
Here’s what may help you have a cleaner code:
Use meaningful variable names that are descriptive and imply type. For example, if you’re declaring a boolean variable about an attribute (age for example) to check whether a person is old, you can make it both descriptive and type-informative by using is_old. The same goes for the way you declare your data: make it explanatory.
# not good ...
import pandas as pd
df = pd.read_csv(path)
# better!
transactions = pd.read_csv(path)Avoid abbreviations that no one but you can understand and long variable names that no one can bear.
Don’t hard code “magic numbers” directly in code. Define them in a variable so that everyone can understand what they refer to.
# not good ...
optimizer = SGD(0.0045, momentum=True)
# better !
learning_rate = 0.0045
optimizer = SGD(learning_rate, momentum=True)Follow PEP8 conventions when naming your objects: for example, functions and methods names are in lowercase and words are separated by an underscore, class names follow the UpperCaseCamelCase convention, constants are fully capitalized, etc. Learn more about these conventions here.
Use indentation and whitespaces to let your code breathe. There are standard conventions such as “using 4 space for each indent”, “separate sections should have additional blank lines”… Since I never remember those, I use a very nice VSCode extension called prettier that automatically reformat my code when pressing ctrl+s.
Source: https://prettier.io/
2 — Make your code modular 📁
When you start building something that you feel can be reused in the same or other projects, you’ll have to organize your code into logical functions and modules. This helps for better organization and maintainability.
For example, you’re working on an NLP project and you may have different processing functions to handle text data (tokenizing, stripping URLs, lemmatizing, etc.). You can put all these units in a python module called text_processing.py and import them from it. Your main program will be way lighter!
These are some good tips I learned about writing modular code:
DRY: Don’t Repeat Yourself. Generalize and consolidate your code whenever possible.
Functions should do one thing. If a function does multiple operations, it becomes more difficult to generalize.
Abstract your logic in functions but without over-engineering it: there’s the slight possibility that you’ll end up with too many modules. Use your judgment, and if you’re inexperienced, have a look at popular GitHub repositories such as scikit-learn and check out their coding style.
3 — Refactor your code 📦
Refactoring aims at reorganizing the internal structure of the code without altering its functionalities. It’s usually done on a working (but still not fully organized) version of the code. It helps de-duplicate functions, reorganize the file structure, and add more abstraction.
To learn more about python refactoring, this article is a great resource.
4 — Make your code efficient ⏱
Writing efficient code that executes fast and consumes less memory and storage is another important skill in software development.
Writing efficient code takes years of experience, but here are some quick tips that may help your find out if your code is running slow and how to boost it:
Before running anything, check the complexity of your algorithm to evaluate its execution time
Check the possible bottlenecks of your script by inspecting the running time of every operation
Avoid for-loops as much as possible and vectorize your operations, especially if you’re using libraries such as NumPy or pandas
Leverage the CPU cores of your machine by using multiprocessing
5 — Use GIT or any other version control system 🔨
In my personal experience, using GIT + Github helped me improve my coding skills and better organize my projects. Since I used it while collaborating with friends and/or colleagues, it made me stick to standards I didn’t obey to in the past.
Source: freecodecamp
There are lots of benefits to using a version control system, be it in data science or software development.
Keeping track of your changes
Rolling back to any previous version of the code
Efficient collaboration between team members via merge and pull requests
Increase of code quality
Code review
Assigning tasks to team members and monitoring their progress over time
Platforms such as Github or Gitlab go even further and provide, among other things, Continuous Integration and Continuous Delivery hooks to automatically build and deploy your projects.
If you’re new to Git I recommend having a look at this tutorial.
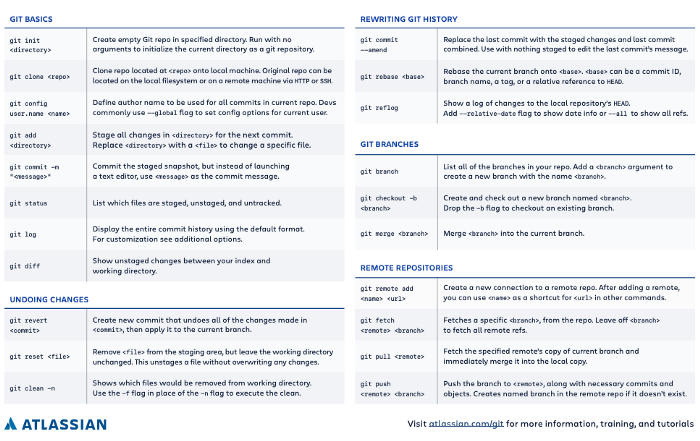
Or you can have a look at this cheat sheet:
source: Atlassian
If you want to specifically learn about how to version machine learning models, have a look at this article.
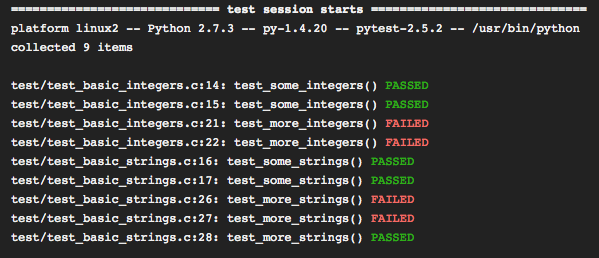
6 — Test your code 📐
If you’re building a data pipeline that executes a series of operations, one way to make sure it performs according to what it’s designed to do, is to write tests that check an expected behavior.
Tests can be as simple as checking an output shape or an expected value returned by a function.
Writing tests for your functions and modules brings many benefits:
It improves the stability of the code and makes mistakes easier to spot
It prevents unexpected outputs
It helps to detect edge cases
It prevents from pushing broken code to production
7 —Use logging 🗞
Once the first version of your code is running, you definitely want to monitor it at every step to understand what happens, track the progress, or spot faulty behavior. Here’s where you can use logging. Here are some tips on efficiently using logging:
Use different levels (debug, info, warning) depending on the nature of the message you want to log
Provide useful information in the logs to help solve the related issues.
import logging
logging.basicConfig(filename='example.log',level=logging.DEBUG)
logging.debug('This message should go to the log file')
logging.info('So should this')
logging.warning('And this, too')source: realpython
References
https://github.com/A2Amir/Software-Engineering-Practices-in-Data-Science.
https://towardsdatascience.com/5-reasons-why-jupyter-notebooks-suck-4dc201e27086
https://medium.com/@_orcaman/jupyter-notebook-is-the-cancer-of-ml-engineering-70b98685ee71
https://datapastry.com/blog/why-i-dont-use-jupyter-notebooks-and-you-shouldnt-either/
https://visualgit.readthedocs.io/en/latest/pages/naming_convention.html
https://towardsdatascience.com/unit-testing-for-data-scientists-dc5e0cd397fb
Conclusion
Long gone the days when data scientists found their way around by producing reports and jupyter notebooks that didn’t communicate in any way with the company systems and infrastructure.
Nowadays, data scientists start producing testable and runnable code that seamlessly integrates with the IT systems. Following software engineering best practices becomes therefore a must. I hope this article gave you an overview of what these best practices are.
Thanks for reading!
Source: paper.li


















Comments